As the smallest and most abundant acellular organisms on Earth, viruses have a profound impact on human health, and play a crucial role in global biogeochemical cycles. Empowered by the emerging metagenomic technologies, a vast number of novel DNA and RNA viruses have been discovered. However, most current virus classification methods primarily rely on nearly complete viral genomic data, resulting in poor classification performance for fragmented and incomplete viral sequences. Furthermore, existing technological approaches tend to work well for classifying double-stranded DNA viruses, whereas accurately classifying RNA viruses and single-stranded DNA viruses in high-throughput settings remains a challenge. Therefore, achieving accurate and detailed classification of environmental viruses has become a crucial challenge that urgently needs to be addressed in the fields of biology, medicine, and ecology. Recently, the research team led by Professor Wang Min from the College of Marine Life developed the VITAP (Viral Taxonomic Assignment Pipeline), a novel viral classification tool, based on sequence alignment and graph theory. There research findings have been published online in Nature Communications. This work provides an important tool for research in viromics and viral ecology.
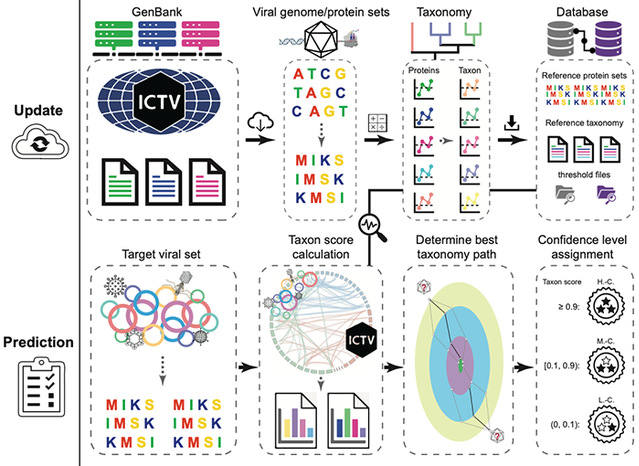
This method combines sequence alignment with graph theory algorithms, which enables accurate classification of both DNA and RNA viruses. It also provides a confidence assessment for each classification unit, and can efficiently classify viral genome sequences down to the genus level, even for sequences as short as 1,000 base pairs, significantly enhancing viral annotation rates. Furthermore, VITAP can automatically update based on the latest classification database established by the International Committee on Taxonomy of Viruses (ICTV), and supports user-defined classification databases (Figure 1).
In the subsequent analyses and tests, the VITAP method was successfully applied to the annotation of metaviromes and viral genomes. The results demonstrated that the accuracy, precision, and recall rates of VITAP all exceeded 0.9 in both family and genus-level classification analyses, and VITAP exhibited a higher annotation rate compared to other methods. In general, VITAP enables a more comprehensive and stable classification of viruses while ensuring high accuracy, thereby providing a new efficient automated tool for research in viral genomics, taxonomy, and ecology (Figure 2).