Eukaryotic gene expression is dynamically regulated through the interplay between histone modifications and chromatin remodeling, yet how these processes are coordinated remains incompletely understood. On February 23, a team led by Professor Gao Shan from the Institute of Evolution and Marine Biodiversity at Ocean University of China (OUC) published a study in Nucleic Acids Research entitled “Bromodomain protein IBD1 bridges histone acetylation and H2A.Z deposition to fine-tune transcription”.
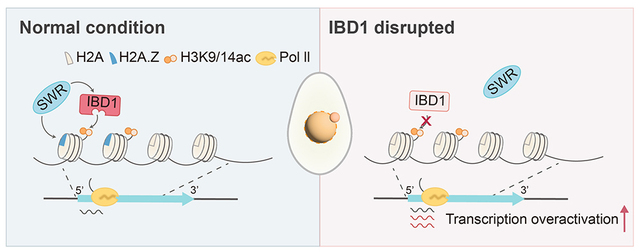
Using the unicellular eukaryotic model organism Tetrahymena thermophila, the study found that IBD1’s bromodomain specifically recognizes H3K9/K14 di-acetylation (H3K9/K14ac) and recruits the SWR complex subunit ARP6, ensuring precise incorporation of the histone variant H2A.Z into chromatin. Perturbation of the histone acetylation-IBD1-H2A.Z regulatory axis leads to widespread transcriptional upregulation. This observation highlights a dual role for H2A.Z: sustaining basal transcription and constraining overactivation at highly active genes. These findings provide new evidence for understanding the biological function of H2A.Z and offer a new perspective on how histone modifications and histone variants cooperate to regulate gene expression.
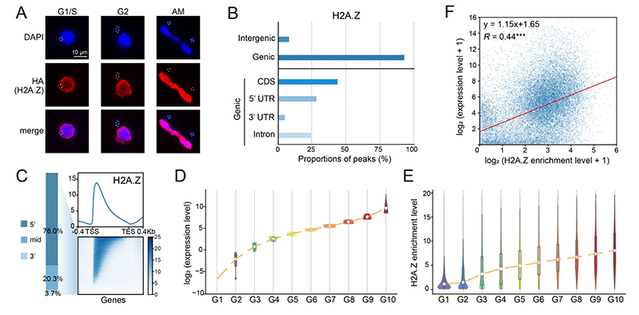
In Tetrahymena thermophila, H2A.Z is exclusively localized in the transcriptionally active macronucleus (MAC), and is enriched at transcriptionally active loci, especially at the 5′ ends of gene bodies. Its enrichment is positively correlated with gene expression levels. These findings support a close association between H2A.Z and transcriptional activity, and provide a foundation for further investigation of its regulatory role.
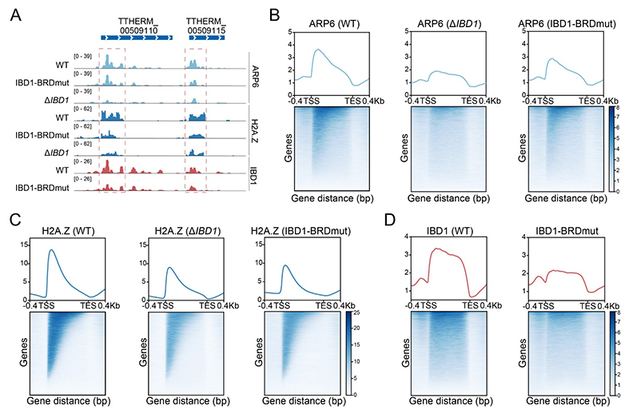
Further sequence and functional analyses showed that IBD1 contains a conserved bromodomain (BRD) capable of recognizing histone acetylation marks. In BRD-mutant (IBD1-BRDmut) cells, IBD1 enrichment at the 5′ ends of gene bodies was significantly reduced; both IBD1 deletion (ΔIBD1) and BRD mutation led to reduced ARP6 enrichment at these regions. As ARP6 is a core subunit of the SWR complex responsible for H2A.Z deposition, these changes further resulted in the reduction of H2A.Z occupancy at the 5′ ends of gene bodies.
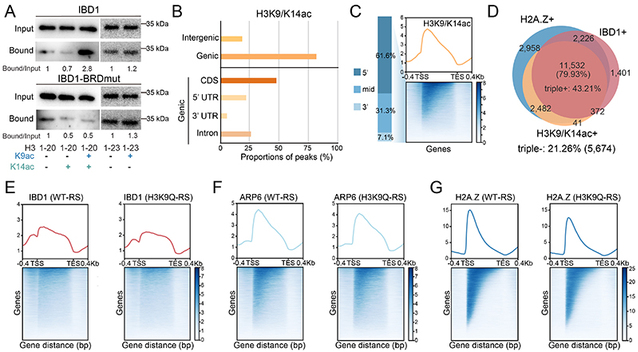
The team then examined the histone acetylation substrate recognized by OBD1 and found that IBD1 preferentially binds H3K9/K14 di-acetylation (H3K9/K14ac). H3K9/K14 and H2A.Z also showed highly similar genomic distribution patterns, with both enriched toward the 5′ ends of gene bodies. In addition, genes enriched for IBD1, H2A.Z, and H3K9/K14ac overlapped extensively. When H3K9 was mutated to Q (H3K9Q-RS), the enrichment of IBD1, ARP6, and H2A.Z at the 5′ ends of gene bodies was reduced.
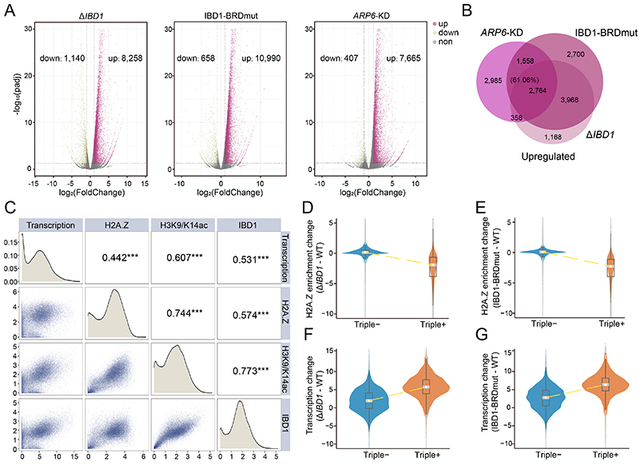
Further analyses showed that impairment of IBD1 function (ΔIBD1 or BRDmut) led to widespread transcriptional upregulation accompanied by an overall reduction of H2A.Z enrichment at the 5′ ends of gene bodies. ARP6 knockdown (ARP6-KD) caused a similar broad increase in transcription. In addition, H2A.Z, IBD1, and H3K9/K14ac were all positively correlated with the expression levels of Pol II-transcribed genes. H2A.Z and H3K9/K14ac enrichment levels were also positively correlated with each other, and both were associated with higher transcriptional activity. Notably, in genes co-enriched for IBD1, H3K9/K14ac, and H2A.Z (triple+), IBD1 deletion or BRD mutation caused a marked reduction in H2A.Z levels together with significant transcriptional upregulation. These effects were more pronounced in highly expressed genes, where H2A.Z depletion was greater and transcriptional upregulation was stronger. Together, these results indicate that H2A.Z is not only positively associated with transcriptional activity but also plays an important role in preventing transcriptional overactivation at highly acetylated genes. At the phenotypic level, this transcriptional dysregulation had severe physiological consequences: both ΔIBD1 and IBD1-BRDmut cells showed markedly reduced growth rates compared with wild-type cells.
Overall, this study establishes a new molecular regulatory pathway in which IBD1 is recruited to chromatin through bromodomain-mediated recognition of histone acetylation, particularly H3K9/K14ac, and then facilitates ARP6-dependent H2A.Z deposition by the SWR complex at the 5′ ends of gene bodies, thereby maintaining transcriptional homeostasis and normal cell growth.








